In recent years, researchers have used DNA to encode everything from Operating system to me Malware. Instead of being a technical curiosity, these efforts were serious attempts to take advantage of DNA’s properties to store data for the long term. DNA can remain chemically stable for hundreds of thousands of years, and we’re unlikely to lose the technology to read it, which is something you can’t say about things like ZIP drives and MO drives.
But so far, writing data into DNA has involved converting the data into a series of bases on a computer, and then arranging that sequence from where a chemical compound acts – organisms don’t actually enter the image. But separately, a group of researchers has been discovering how to record biological events by modifying a cell’s DNA, allowing them to read the cell’s history. A group at Columbia University has now figured out how to combine the strains and write the data into DNA using voltage differences applied to live bacteria.
CRISPR and data storage
The CRISPR system was developed as a way to completely edit or cut genes from DNA. But the system caught biologists for the first time because it introduced new sequences into DNA. For all details, see Cover our NobleFor now, however, you only need to know that part of the CRISPR system involves identifying the DNA from viruses and inserting copies of it into the bacterial genome in order to identify it if the virus appears again.
The group in Colombia figured out how to use this to record memories in bacteria. Suppose you have a process that activates genes in response to a specific chemical, such as sugar. The researchers also switched this to activate a system that makes copies of a circular piece of DNA called a plasmid. Once the build number went up, they activated the CRISPR system. Given the circumstances, it was more likely that a copy of the plasmid DNA was introduced into the genome. When sugar is not present, it generally enters something else.
With this system, it was possible to find out if the bacteria had been exposed to sugar in the past. It’s not perfect, because the CRISPR system doesn’t always pop something in when you want it, but it does run average. Therefore, you only have to arrange a sufficient number of bacteria in order to know the average sequence of events.
To adapt this for data storage, the researchers used two plasmids. One is the same as shown above: it is present at low levels when a particular signal is absent, and present at very high levels when the signal is present. The second is always present at moderate levels. When CRISPR is activated, it tends to introduce sequences of the plasmid present at higher levels, as shown in the diagram below.
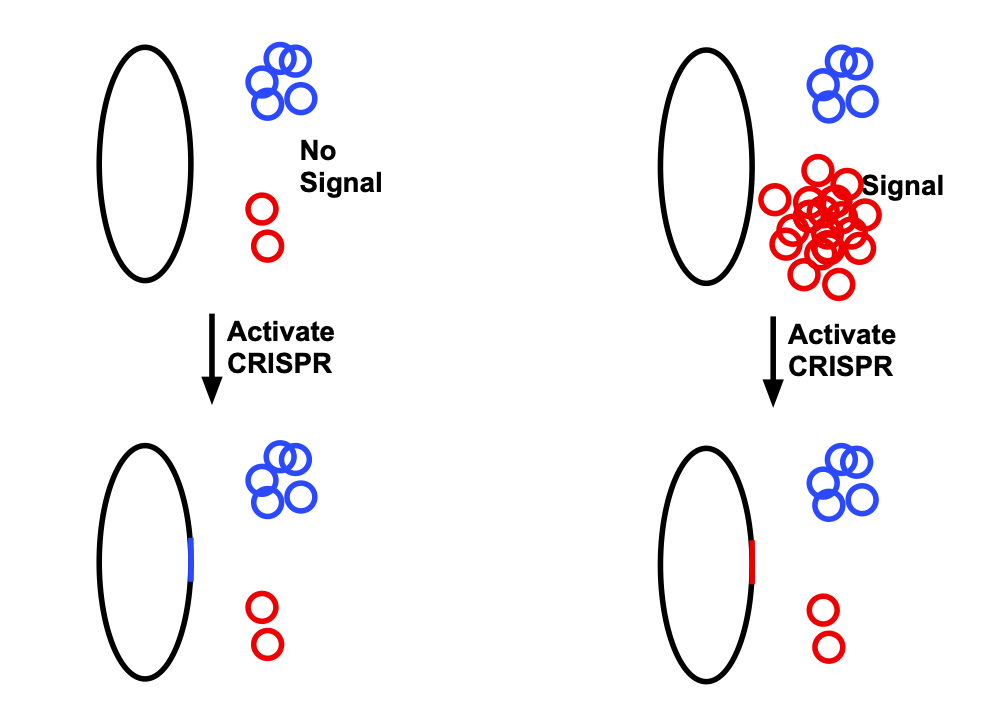
John Timmer
On its own, this stores only one bit. But the process can be repeated, creating an extension of DNA that is a series of insertions derived from red and blue plasmids, with identification determined by whether or not the signal is present.
Give her a shake
It’s a neat system but it’s a long way from the kinds of things we usually associate with producing data – rarely the product of a sensor reading or calculation is a sugar or an antibiotic mixed with a group of bacteria. It turns out that getting bacteria to respond to an electrical signal is relatively simple. Coli Able to change gene activity depending on whether it is in an oxidizing or reducing chemical environment. And researchers can alter the environment by applying voltage differences to a specific chemical on the farm with the bacteria.
More specifically, the potential difference alters the oxidative state of a chemical called ferrocyanide. This, in turn, caused the bacteria to alter gene activity. By engineering the plasmid so that it responds to the same signal as these genes, the researchers were able to control plasmid levels by applying a different electrical potential. They can then record this level of this plasmid by activating the CRISPR system in these cells.
It’s very easy to see how each of the entries in a string can be considered either zero or one, depending on the identity of the extension. But remember that this system is not perfect; Quite regularly, the CRISPR technology won’t input anything when energized, which can displace all subsequent bits. Since this process is random, the longer the string of bits you are trying to encode, the more likely it is that at least one of them will be skipped.
To reduce this problem, the researchers kept their data to three bits per group of bacteria. Until then, they had to train a moderated learning algorithm to reconstruct the most probability bit strings based on the average of sequences present in the population. Even so, the system failed to recognize the bit string about six percent of the time. In the end, they settled on using the parity bit which was the sum of the first two to allow for error correction, and then they edited a lot of groups in parallel.
(By giving each group a unique sequence marker called a “barcode” plasmids, it was possible to mix many of them into one group after encoding the bits and continuing to decode everything once the DNA had been sequenced).
With everything in place, they stored and read “Hello world!” They even put bacteria in some cultivated soil for a week and showed that they are able to recover the message. (Storing it in the freezer obviously works best.) They estimate that the message can be kept for as many as 80 generations of bacteria.
Let’s be clear: As a storage medium, in its current form, this is pretty appalling. If you were to put some data in DNA, it would be much better if the DNA was chemically synthesized. But it is interesting to think that we can go directly from the electrical signals to the altered DNA, and there may be some ways to improve the system now that it is built.
Biology of Chemical NatureDOI: 2021. 10.1038 / s41589-020-00711-4 (About DOIs).

“Appassionato di alcol. Piantagrane. Introverso. Studente. Amante dei social media. Ninja del web. Fan del bacon. Lettore”.